RNAi Analysis Using Near-Infrared Fluorescence
RNAi Analysis Using Western Blots
Quantitative Western blotting is an essential tool for every step in your RNAi studies. With quantitative Western blots, you can:
- Choose an RNAi vehicle (siRNA or shRNA or other RNA-producing vector construct)
- Confirm level of knockdown of the target gene
- Determine the impact of target gene knockdown on a phenotype such as cell proliferation, cell migration, cell cycle, or cell signaling pathways
The Odyssey® M and Odyssey DLx Infrared Imagers provide accurate digital protein data for your RNAi studies.
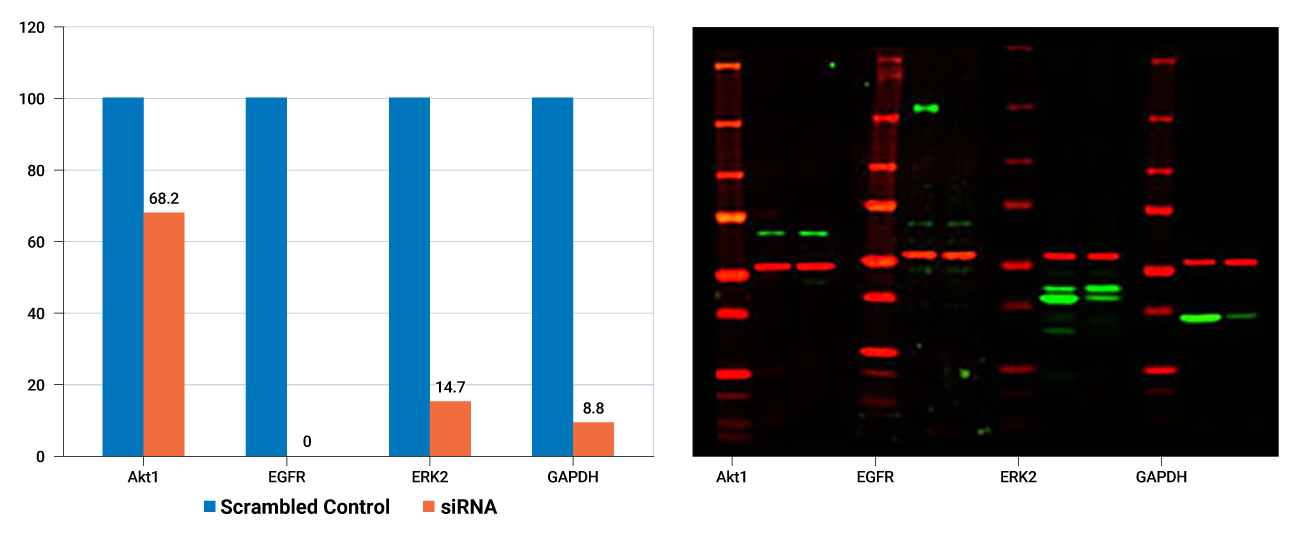
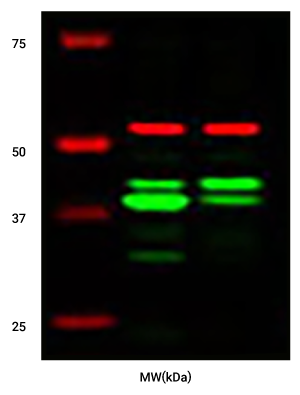
- Raw data can be normalized for increased accuracy using one of the two independent channels (colors) to monitor a control protein.
- Stable fluorescent signal is directly proportional to amount of target for consistent data.
- Wide linear range allows knockdown levels involving both weak and strong signals to be measured accurately in a single image without the bleed over of signal from adjacent "blow out" bands.
RNAi Analysis Using In-Cell Western™ Assays
The In-Cell Western Assay can be used in a functional siRNA screen measuring the effects of knockdowns in cultured cells.
In-Cell Western assays offer additional throughput for more complex studies, as well as exceptionally consistent data (Z'-factor).1 The ICW has been demonstrated as a powerful cellular assay for genome-wide RNAi screens.2
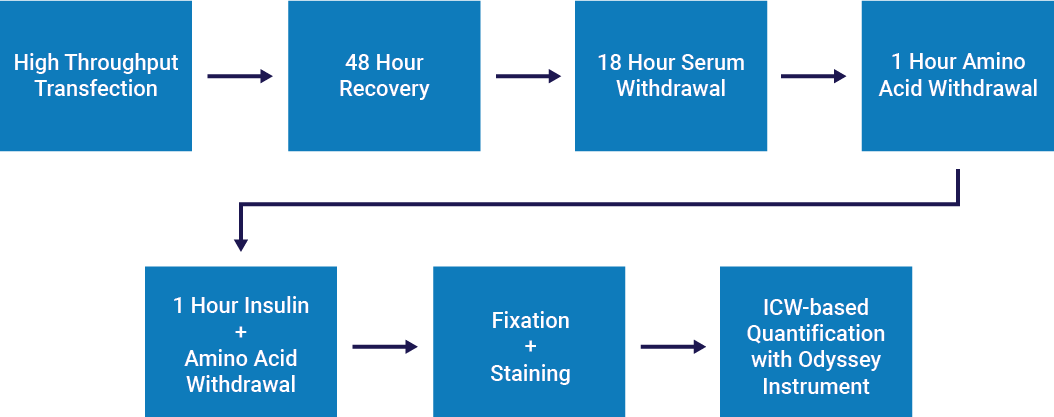
Hoffmann et al.2 used In-Cell Western screening to assess the effects of knockdowns on mTORC1-dependent phosphorylation of ribosomal protein S6 (rpS6).
- HeLa cells were transfected with siRNA and screened for phosphorylation of prS6 at Ser235/236.
- NHS ester cell labeling was used for cell number normalization.
- The Dharmacon Human Druggable Genome siRNA library gave 7,317 genes from the human genome that are likely targets for pharmacological inhibition.
- A pilot small molecule screen was performed with a library of ~2500 compounds.
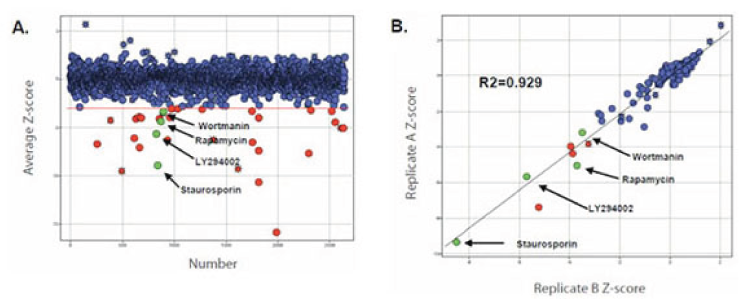
- Knockdown of components required for both the growth factor and amino acid branches of the mTORC1 signaling network caused reduction in phospho-rpS6.
- Known genes involved in growth factor-mediated inputs leading to rpS6 phosphorylation (IRS2, IGF1R, PI3-kinase, PDK1, and S6K2) scored as hits in the screen, validating the approach.
- In addition to known pathway components, several uncharacterized genes scored as strong hits.
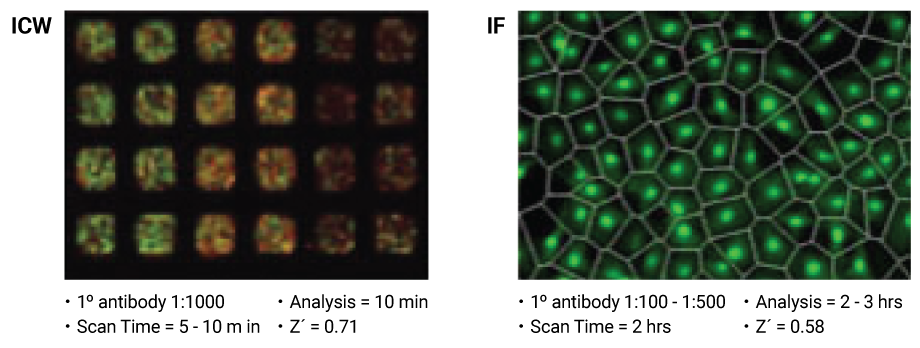
In-Cell Western RNAi screening was found to be faster and less expensive than high content microscopy (IF), and gave similar or better statistical replicability.
Resources
References
- Boveia, V et al. (2009). Using the Z’-Factor Coefficient to Monitor Quality of Near-Infrared Fluorescent Cell-Based Assays. LI-COR Biosciences.
- Hoffmann et al. (2008). A functional siRNA screen for novel regulators of mTORC1 signaling. Poster presentation, ASCB Annual Meeting.