Optical Probe Development Applications
Imaging in the NIR Spectrum
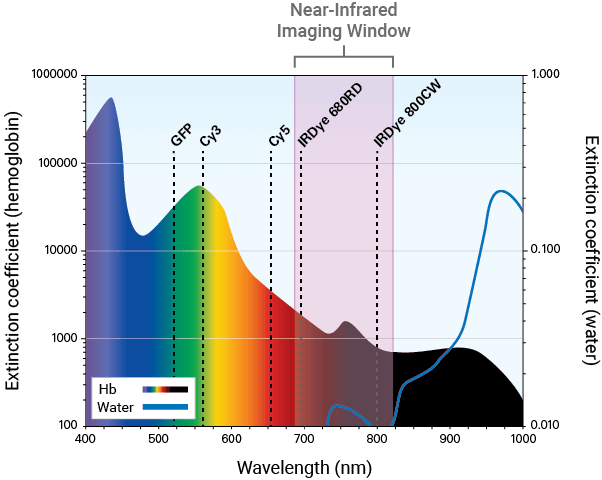
Near-infrared fluorescence detection (680 nm to 820 nm) is a sensitive and efficient modality for in vivo studies. NIR optical imaging provides several advantages, such as:
Sensitivity
Light absorbance by, and background autofluorescence from animal tissue is significantly lower in the NIR region. These properties allow better light penetration and higher signal-to-noise ratios for detecting deeper targets in vivo.
Throughput
In contrast to other imaging modalities, images can be captured within seconds, allowing rapid screening of mice.
Cost
Compared to other modalities, optical imaging is more economical.
These characteristics make NIR detection ideal for a variety of in vivo imaging applications.
"The Pearl is very easy to use, and it's been very helpful for our optical imaging agent development program."
Optical Probes for NIR Imaging
Optical probes aid in visualizing and monitoring the pharmacology of the targeting agent of interest. Near-infrared labeled targeting agents include antibodies, proteins, peptides, ligands, and drugs, among others.
Developing an optical probe for diagnostic or therapeutic applications involves several validation steps. Characterization of optical probes should be done in vitro to confirm specificity to the target before moving to animal studies, as in vivo studies can be very costly. Biological characterization of the optical probe in vivo and ex vivo can follow using the Pearl® Trilogy Small Animal Imaging System.
Target Specificity Studies
Why Assess Target Specificity?
Target specificity studies evaluate the binding specificity and affinity of the probe to targets in vivo. These studies also help identify any non-specific interactions of the probe. Cell surface receptors and transporters are the most common targets for in vivo specificity studies.
Receptor Targeting
Cell surface receptors such as EGF and integrin receptors are commonly targeted for in vivo imaging in cancer. Tumors often over-express cell surface receptors, providing easy access for targeting molecules such as ligands, antibodies, or drugs. These targeting molecules bind to the receptors, and thus can be evaluated for imaging studies.
Transporter Targeting
For in vivo imaging of cancer, cell surface transporters such as glucose transporters are also commonly employed as targets. This is because tumors are often more metabolically active than surrounding normal cells, and may have an increased rate of glucose metabolism and cell surface glucose transporter activity.1, 6 For instance, PET imaging employs the binding property of FDG to GLUT transporters in imaging.
Evaluating Target Specificity using NIR Fluorescence
Low autofluorescence from tissue components and deeper penetration of light in the near-infrared spectrum allows sensitive detection and semi-quantitative assessment of probe binding affinity to the intended target. Non-specific interactions can also be detected with high sensitivity. Additionally, uniform laser illumination across the entire imaging area in the Pearl Trilogy enables accurate in vivo measurement of specificity.
Probes can be developed against several receptor and transporter targets. For instance, the GLUT1 glucose transporter can be targeted with a fluorescent-labeled IRDye® 2-deoxyglucose (2-DG) agent for longitudinal studies of tumor progression. 2 EGF and integrin receptors can be targeted using IRDye EGF agent 3 or IRDye 800CW RGD optical probes,4, 5 respectively.
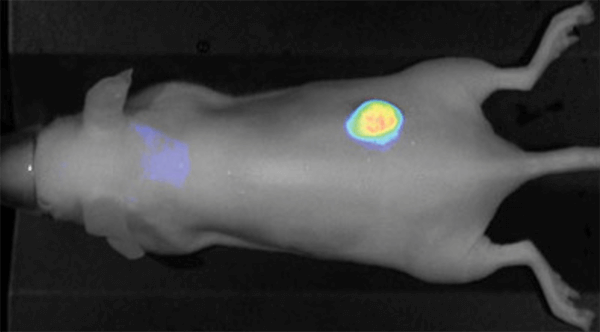
Figure 1. IRDye 800CW EGF imaged 96 h post injection. Image was captured with the Pearl Imager with the pseudo-color representing the signal in the 800 nm channel overlaid on the mouse white light image.
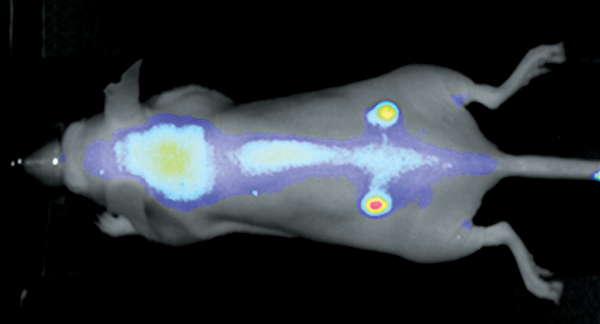
Figure 2. Nude mouse bearing subcutaneous tumors, U87 (left hip) and A431 (right hip), was imaged 24 h post intravenous injection of IRDye 800CW RGD (1 nmole). Image was captured on the Pearl Imager; 800nm signal is presented in pseudo-color overlaid on a white light image of the mouse.
Biodistribution and Clearance Studies
Why Measure Biodistribution and Clearance?
Biodistribution and clearance assessments track the movement of a probe in order to understand its uptake, localization, distribution, and elimination within the animal.
Biodistribution studies can identify regions and time of specific and non-specific retention of the probe in organs and tissues.
- In vivo biodistribution analysis in live animals tracks the localization of the probe and reveals if it binds to the intended target, as well as its elimination route. Non-specific interactions of the probe are very important and should not be overlooked.
- For ex vivo biodistribution analysis, tissues excised from the animal (tumors, organs) are assessed for relative accumulation of the probe. Macroscopic analysis of tissue sections and whole organ imaging allow a quick screening overview, while microscopic analysis provides additional detail of cellular localization of the probe.
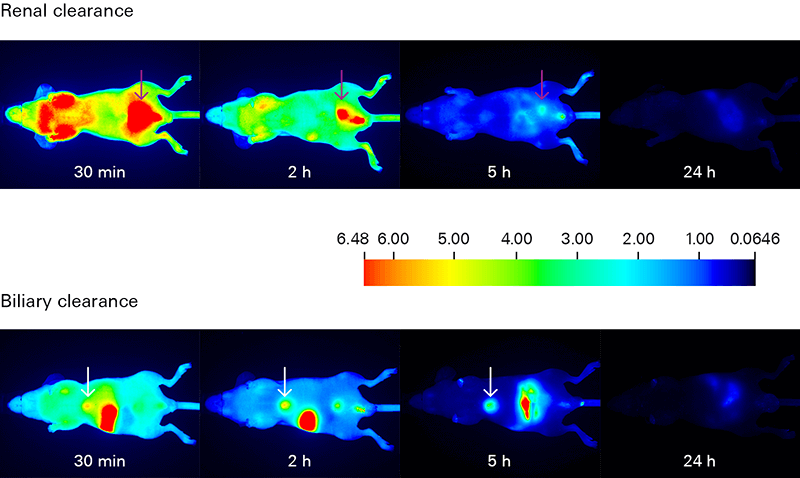
In vivo clearance studies monitor the elimination of the optical probe over time, in live animals. Knowledge of probe elimination is important to optimize both imaging time and dose, as well as to identify any specific or non-specific retention, which may impact imaging results.
Assessing Biodistribution and Clearance
using NIR Fluorescence
Deeper light penetration and low autofluorescence from animal tissues make near-infrared detection ideal for tracking biodistribution (in vivo and ex vivo) and clearance with high sensitivity. Low background yields excellent signal-to-noise ratios for monitoring movement of probes in vivo.
The Pearl Trilogy Imaging System can perform rapid time-lapse and longitudinal in vivo NIR imaging without the need to manually adjust camera settings between scans. For ex vivo analysis, the Organ Tray Base and disposable organ trays provide a non-heated, flat surface for imaging on the Pearl Trilogy system. IRDye infrared dyes IRDye 800CW and IRDye 680RD provide excellent performance for in vivo and ex vivo biodistribution imaging.
References
- Gambhir, SS et al. (2001) A Tabulated Summary of the FDG PET Literature. Journal of Nuclear Medicine 42(5), pp 1S-93S.
- Kovar, J et al. (2009). Analytical Biochemistry, 384(2), pp 254-62. DOI: 10.1016/j.ab.2008.09.050.
- Kovar, J et al. (2006). Am J Pathol, 169(4), pp 1415-26. DOI: 10.2353/ajpath.2006.060324.
- Kovar, J et al. (2009). Poster presentated at AACR Annual Meeting.
- Chen, K et al. (2009). Molecular Imaging, 8(2), pp 65-73. DOI: 10.2310/7290.2009.00011.
- Warburg O. (1956). Science, 123(3191), pp 309-14. DOI: 10.1126/science.123.3191.309.
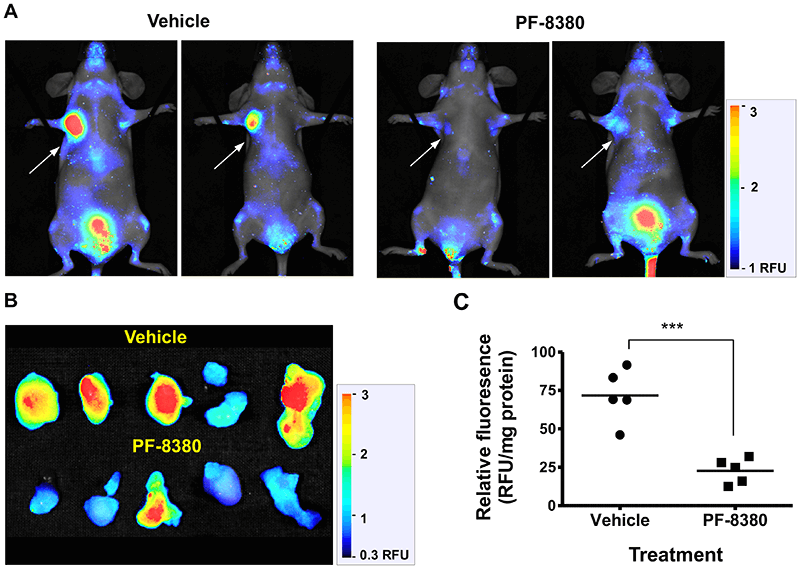
- Madan D, et al. (2013). PLOS ONE, 8(11). DOI: 10.1371/journal.pone.0079065.
